Nowadays, we are witnessing an unprecedented revolution in life sciences technologies, like the one experienced in physics and electronics during the past century. Once these living technologies will reach maturity, there will be an impact in our society, complementing, or even replacing some of the current electro-mechanic technologies. In this context, there is a clear consensus that the development of living technologies compulsorily requires the development of complex control systems that integrate multiple inputs and respond according to complex decisions.
Taking inspiration from already existing methodologies, we looked at the major progress being made on printed electronics. By having a set of inks with metal particles, it is possible to conduct electricity in a large variety of substrates, generating great demand, as smart functions can be integrated to existing products.
Having this idea of patterning and having great experience in the field of Synthetic Biology, we came up with the methodology proposed in the article “2D printed multicellular devices performing spatial computation for digital and analogue circuits”.
In this study, we propose a method to construct printable cellular devices, digital or analogue, for different potential purposes, such as complex diagnostics implying different input logic integration.
These printable devices are designed to work in a 2D surface, in which the circuit information is encoded in the concentration of a biological signal, the so-called carrying signal (i.e. like electric current in electronics). This signal diffuses through the 2D surface and thereby interacts with different device components. These components are distributed in a specific spatial arrangement and perform the computation by modulating the level of the carrying signal in response to external inputs, determining the final output.
These 2D cellular devices have been printed on paper surfaces, (even though other materials have been explored, such as nylon) using a set of cellular inks. As a proof-of-principle, we have printed and analysed both digital and analogue circuits using the same set of cellular inks but with different spatial topologies.
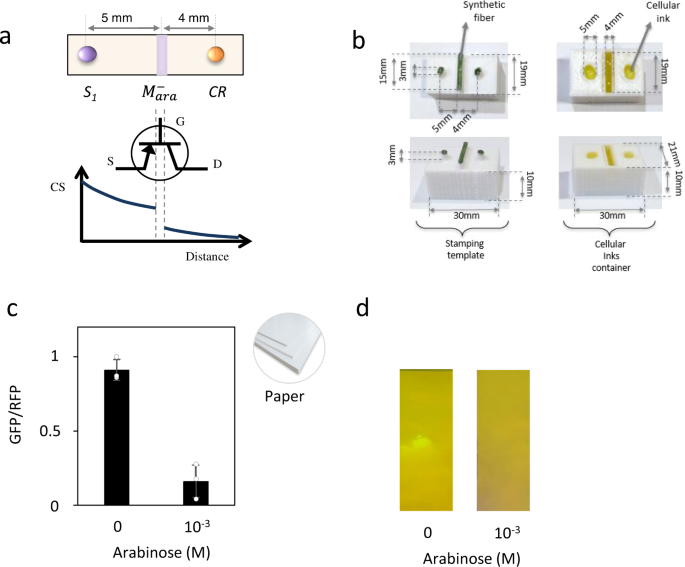
To validate this methodology, we have built a library of engineered cells, each of them accounting for a different circuit element. Here we show a schematic figure to explain the system logic. As seen in Figure 1a, a living transistor-like circuit can be implemented, having a biological source (i.e. AHL molecules), a gate composed of two different biological modulators which can allow or block the flow of our biological signal depending on external inputs, and a drain which generates the final output, in this case, a cell producing or not fluorescence (GFP). Figure 1b shows the stamping system we have used for circuit automatization. Stamping offers numerous advantages over previous methods for cell printing such as 3D printers. For instance, stamping is faster than 3D printing, does not require complex printing devices, and templates can be easily customized, which makes stamping suitable for industrial production, potentially at very low cost and with minimal requirements.

Figure 1. Printed circuit implementing a transistor-like device. a. Mapping of a general transistor architecture on a cellular printed pattern. The cellular pattern is composed of three main components: source (S1 cells), gate (M- cells) that responds to external inputs and a drain (CR cells) as the final output responding to the presence of the carrying signal (CS). b. Stamping system. Containers for circuit design. c. Circuit response. In the absence of external input, i.e. arabinose, the CS encoded in the production of AHL molecules by S1 cells diffuses along the surface, inducing GFP expression in reporter cells CR. In the presence of 10-3 M arabinose (Ara), the modulatory element M-ara produces the AHL cleaving enzyme Aiia, which degrades the CS. d. Picture of the device response.
From this first approach, we have established a general architecture to translate any arbitrary logic circuit into a 2D living device and we have been able to develop more complex devices.
By playing with distributed computation and spatial segregation we have been able to overcome some constraints regarding the wiring problem or the genetic complexity of current circuit designs, which make them unsuitable to be used out of the lab.
Overall, we believe printable cellular circuits could represent an interesting alternative for paper-based devices with appealing potential advantages, such the possibility of producing low-cost cellular devices based on a simple and autonomous experimental setup methodology, easing their technological transference. Further development is required to explore this potential and meet future market needs.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in