In the late summer of 2018, I met Dr. Haoyu Li from the Harbin Institute of Technology. At that time, he had just finished his postdoc training in the States and started to build his lab. Through the introduction from a mutual friend, we invited him to give a talk on his work. I still remember the first impression, how he enthusiastically presented the new algorithm to improve the resolutions of images obtained from a light-field microscope. Given those light-field microscopes suffered from limited and heterogenous spatial resolutions along with different focal positions, I wondered whether our structure illumination microscopy (SIM) data could be a better example to demonstrate the power of their algorithm.
At that time, we had just published our first Nature Biotechnology paper on Hessian structured illumination microscope (Hessian-SIM)1. Using the spatiotemporal continuity of any signals as the general prior knowledge to reduce intrinsic artifacts underlying the Weiner reconstruction, we have realized SIM imaging at ~90 nm, 564 Hz (by rolling reconstruction) in cells labeled with standard fluorescent probes. Despite this unprecedented combination of spatiotemporal resolution, we were still asked by other researchers the question all the time: can we push the spatial resolution of live-cell superresolution (SR) further without much sacrificing of other parameters? Therefore, although I was skeptical whether that was a feasible goal myself, we are interested in any algorithms that may help resolve intricate and dynamic structures in live cells with a limited photon budget and stringent requirements of photo-toxicity.
The sparse deconvolution algorithm is a two-step protocol. First, we have modified the previous iterative optimization procedure by adding the relative sparsity constraint on top of the continuity one. Next, we used a Richardson-Lucy deconvolution2, 3 algorithm to process the fluorescence image and push the resolution limit further. After deconvolution with this algorithm, much better contrast images of mitochondrial cristae structures (labeled with Mito-tracker Green, Fig.1) and actin meshes (labeled with lifeact-EGFP) throughout the cell emerged. However, I wondered whether the visual effects of increased resolvability were due to extension of resolution out-of-the optical transfer function (OTF) support or increase of contrast within the high-frequency domain of the OTF.

Fig. 1 | Relative movements between sub-organellar structures observed by dual-color Sparse-SIM. (f) A representative example of both the IMM (cyan) and ER (magenta, Sec61β-mCherry). As shown in the inset, we found that ER tubules randomly contacted the mitochondria with equal probability at both the cristae regions and the regions between cristae. (g) Magnified views from the white box in (f). We found that the contact between one ER tubule and the top of a mitochondrion not only correlated with the directional movement of the latter but also the synergistically rearranged orientations of cristae thereafter (Supplementary Video 8). Scale bars: (f) 1 μm; (g) 500 nm.
Indeed I told Haoyu and the major contributing student, Weisong Zhao. "Let's keep our noses down and focus on the increase of the contrast and the removal of out-of-focal plane haze by the sparse deconvolution will. Talking about further increasing the resolution out of OTF support is challenging to prove, and nobody will believe us." They agreed, and thus we tried the algorithm on fluorescence images of different structures and microscopes. So it was a surprise to me to see simulated paired lines 60 nm apart well separated after the sparse deconvolution (Fig. 2).

Fig. 2 | Benchmark of spatial resolution at different steps of sparse deconvolution according to the synthetic images. (a) The resolution plate with a pixel size of 16.25 nm, which contained five pairs of lines at distances of 48.75 nm, 65.00 nm, 81.25 nm, 97.50 nm, and 113.75 nm. The synthetic image (512 × 512 pixels) is first illuminated by pattern excitation and then convolved with a microscope PSF of (1.4 NA, 488 nm excitation). The signal is recorded with an sCMOS camera with a pixel size of 65 nm, which meant 4× downsampling of the original image (128×128 pixels). We also included Gaussian noise with a variance of 5% of the peak intensity of the line to the raw image. Next (from left to right), we used inverse Wiener filtering to obtain a conventional SIM image (256 × 256 pixels), followed by the reconstruction constrained by continuity and sparsity a priori information and final deconvolution. The theoretical limit resolution of Wiener SIM was calculated to be 97.6 nm by following the equation λ/2(NAi+NAd), in which i and d respectively represent the illumination and detection NA.
"In simulations, anything could happen." I insisted. "You must show me hard-core evidence. For example, previously ring-shaped caveolae structures were only shown in live cells using nonlinear SIM. If the system resolution could really be improved, we shall be able to visualize that structures." So Shiqun Zhao and Liuju Li went on to do the experiment, while Weisong processed the data once data were available. When they came along with the data, it was shocking to see that we indeed saw small ring-shaped structures (Fig. 3). At that moment, I began to think of the possibility of OTF extension seriously. 
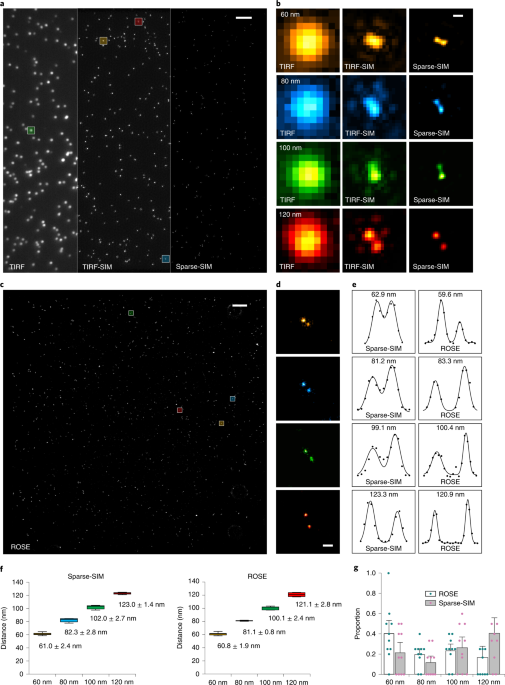
Fig. 3 | Sparse-SIM achieves ~60-nm and millisecond spatiotemporal resolution in live cells. j, A representative COS-7 cell labeled with caveolin–EGFP. k,l, From top to bottom are magnified views of the white box in j reconstructed by TIRF-SIM, Sparse-SIM and Sparse-SIM ×2 with upsampling (k), and their fluorescence profiles are shown in l.
Digging deep into old piles of literature, we realized that another class of thoughts had been around for a long time. Analogous to communication systems, optical systems are channels that transmit information from the object plane to the image plane. In that sense, the spatial bandwidth of the optical system traditionally regarded as invariant may not be the fundamental limit; instead, the ultimate limit may be set by the degrees of freedom (the minimal number of independent variables needed to describe an optical signal) that the optical system can transmit. Wolter proposed mathematical bandwidth extrapolation in objects with finite-size4. As nicely summarized in Goodman's book5, the fundamental principle is that the two-dimensional Fourier transform of a spatially bound function is an analytic function in the spatial frequency domain6. If an analytic function in the spatial frequency domain is known exactly over a finite interval, it has been proven that the entire function can be found uniquely using analytic continuation6. For an optical system, it follows that if an object has a finite size, a unique analytic function exists that coincides inside G(u, v). By extrapolating the observed spectrum using algorithms such as the Gerchberg algorithm7, it is possible in principle to reconstruct the object with arbitrary precision. By making some assumptions about the image and object statistics, the iterative Richardson–Lucy (RL) deconvolution2, 3 represented another class of computational SR methods, which could surpass the Rayleigh criterion in separating double stars in astronomical imaging8. However, such astronomical SR imaging was infeasible for solar systems9. Finally, people proposed a compressive sensing paradigm that enables SR in proof-of-principle experiments but failed in actual applications10. Stable reconstruction always depends critically on the accuracy and availability of the assumed a priori knowledge5, 6, 11 and logarithmically on the image signal-to-noise ratio (SNR)10, 12, 13. How these different bandwidth extrapolation methods reconcile remain unknown. Most importantly, despite the theoretical feasibility, it is generally agreed that the Rayleigh diffraction limit represents a practical frontier that cannot be overcome by applying bandwidth extrapolating methods on images obtained from conventional imaging systems5.

Fig. 4 | Low-pass filtered image recovered by RL deconvolution under noise-free condition. (a) The ground truth contains synthetic various-shaped structures with a pixel size of 20 nm. (b) The low-pass filtered (equal to 90 nm resolution) image of (a). (c) RL deconvolution result of (b) with 2×104 iterations. (d) RL deconvolution with 2×107 iterations. (e-h) Fourier transforms of (a-d). (i-l) Magnified views of white boxes from (a-d). (m) The corresponding profiles are taken from the regions between the yellow arrowheads in (i-l).
In contrast to previous attempts, the success of our application in improving resolutions of different fluorescence microscopes may lie in the heart of two prior knowledge used. The continuity a prior is critical to suppress noise and boost SNR, while the sparsity a prior may create the relatively sparse samples for the later RL deconvolution. As we have shown, the blurred and OTF-filtered structures without noise can be nearly perfectly restored to their original shapes by numerous rounds of RL deconvolution (Fig. 4). Therefore, from raw images, the first step may create the ideal fluorescence images for the later stage of iterative RL deconvolution, and both two components are required for the algorithm to perform the best.
Having speculated the potential mechanism for resolution extension, we excitedly submitted the whole package to prestigious journals at the beginning of 2020. Then we met with an avalanche of doubts regarding the validity, accuracy, and adaptability of our method and data over and over again. In short, I listed the most critical questions below:
- How does the method differ from previous deconvolution methods, especially our previous Hessian deconvolution?
- Is this an actual increase in resolution or just results of oversharpened data or over-clipped background?
- How shall you adjust the continuity and sparsity parameters in the software to obtain the ideal results?
- Can one set of parameters fit structures of different sizes and shapes?
- The results of sparse deconvolution need to be benchmarked with known structures or results obtained by other SR imaging modalities.
- Based on what prior knowledge do you choose the ideal set of parameters?
From these questions, we can see that people are skeptical regarding our general bandwidth extrapolation method. After all, when the OTF cuts off the high-frequency information of the microscope, it seems impossible to retrieve the lost information back, as there are theoretically infinite choices for structures beyond OTF limits. To persuade reviewers and other people, we have worked very hard for the next one and a half years to address each and every one of these questions. Subsequently, we added much more confirmatory data and analysis in the supplementary, provided step-by-step guidelines for the protocol of parameters adjustments, and listed the limitations we found in improving resolution and/or contrast.
Among all questions, Question 6 is the most penetrative one that leads us to a philosophical dilemma: "You must assume an a priori expectation of what you 'should' see in the data to choose the idea parameters and avoid artifacts. However, if you already know the structure in advance, then the method may not (fatally) be so useful. "When I saw this comment, I was stuck and depressed for several days because it does seem to be the precise strike on the crack in our armor. It was fortunate for me to come across two papers written by Teng-Leong Chew and colleagues14 in the Journal of Cell Science, in which they advised how we should accurately report in image acquisition and processing for others to replicate and reproduce the data and analysis. In the paper, I notice that people have used visual interpretation to determine ideal iteration numbers of deconvolution all the time. So we are not alone in finding guilty in this aspect.
Based on that perception, we went a step further, and tried to catch the essence underlying "good" results conferred by the visual inspection. We conclude that sparsity and continuity are again the prior knowledge needed underlying such judgments. For example, while increasing the sparsity value, we looked for the emergence of additional high-frequency structural information allowed by the Nyquist sampling criteria of the designated resolution from deconvolved images. Although higher resolution may be achievable in images with superior SNR, we selected 60 nm as the usual resolution limit for the Sparse SIM to avoid uncertainties associated with over-interfering. For images with a pixel size of ~32 nm, appearances of structures (lines, puncta) with the FWHM less than 2 pixels (64 nm) were often regarded as "over filtering" artifacts unless more specific prior knowledge is assumed. Otherwise, we advised to turn down the sparsity value. In contrast, sparse deconvolution reconstructed structures larger than or similar in size to those in the original image were due to the over-smooth effects that required down-adjusting the fidelity value.
One last thing to note, different sets of optimal parameters lead to similar but not identical images, (Fig. 5), which may complicate quantitative image analysis. However, this is not new since "image processing is a critical step of almost any microscopy-based experiment, the chosen details of an image processing workflow can dramatically determine the outcome of subsequent analyses"14. Therefore, some personal-biased prior knowledge may lead to different fluorescence intensities of the same viral particle estimated as no ground truth is known. However, as a "deterministic, much more mathematically tractable" method, our sparse deconvolution can be repeated if detailed parameters are given, which is superior to the popular machine learning techniques in this perspective. Thus, as Teng-Leong Chew and colleagues proposed14, we urged users to keep the original and deconvolved images and documentation of all parameters. Biological researchers can reproduce the process or adjust parameters to present and re-analyze the data better. On the other hand, technology developers can rigorously explore our codes and procedures to improve the method further in the future.

Fig. 5 | Partial ring/line simulation. (a) We created synthetic partial and complete ring structures with an 80 nm diameter of 120 nm, 80 nm, 40 nm, and 0 nm apart as ground-truth. (b, c) The ground-truth image in (a) was convolved with a PSF with FWHM of either 110 nm (b) or 45 nm (c), and such images were subsequently subsampled 16 times to obtain the pixel size of 16 nm, and corrupted with Poisson noise and 5% Gaussian noise. (d) The images in (b) were reconstructed by LW deconvolution without sparsity and continuity constraints. (e, f) The images in (b) were reconstructed by Sparse deconvolution under different parameters. The fidelity and sparsity values were set as 300 and 40 for results in (e), and the results for 1200 and 115 parameter conditions were shown in (f).
Looking back in time, moving into this field was quite an accident, and the journey to convince ourselves and other people has been challenging and rocky. Right at this moment, many questions remain unresolved since our knowledge only slowly expands after many endowers. For example, we do not have a clear, mechanistic picture of how the method works, nor do we fully understand its boundary and limitations. The searching for optimal parameters is still manual, tedious, and challenging. However, we argue that our sparse deconvolution confers a general computational method to improve the ability of fluorescence microscopes' ability to resolve structures beyond those permitted by their hardware optics, thus shall be of high potential impact for the life sciences researchers. By providing sufficient experiments and documentation to demonstrate and ensure the usefulness, validity, transparency, and reproducibility of the sparse deconvolution, it is time for people themselves to explore its boundary and potential. This work also constitutes a decisive step into the first practical computational SR method. It suggests that the fundamental limit may be the degrees of freedom the microscope can transmit, but not the spatial bandwidth. Hopefully, by exploring the a priori knowledge of the object, more researchers will continue to push the microscope spatiotemporal resolution limit in the future.
References
- Huang, X. et al. Fast, long-term, super-resolution imaging with Hessian structured illumination microscopy. Nature biotechnology 36, 451-459 (2018).
- Richardson, W.H. Bayesian-based iterative method of image restoration. Journal of The Optical Society of America A 62, 55-59 (1972).
- Lucy, L.B. An iterative technique for the rectification of observed distributions. The Astronomical Journal 79, 745 (1974).
- Wolter, H. On basic analogies and principal differences between optical and electronic information, Vol. 1. (Elsevier, 1961).
- Goodman & W, J. Introduction to Fourier optics. (Roberts and Company Publishers, 2005).
- Bertero, M. & De Mol, C. Super-Resolution by Data Inversion, Vol. 36. (Elsevier, 1996).
- Hunt, B.R. Super-resolution of images: Algorithms, principles, performance. International Journal of Imaging Systems and Technology 6, 297-304 (1995).
- Lucy, L.B. Resolution limits for deconvolved images. The Astronomical Journal 104, 1260-1265 (1992).
- Puschmann, K.G. & Kneer, F. On super-resolution in astronomical imaging. Astronomy & Astrophysics 436, 373-378 (2005).
- Gazit, S., Szameit, A., Eldar, Y.C. & Segev, M. Super-resolution and reconstruction of sparse sub-wavelength images. Optics Express 17, 23920-23946 (2009).
- Lindberg, J. Mathematical concepts of optical superresolution. Journal of Optics 14, 083001 (2012).
- Demanet, L. & Nguyen, N. The recoverability limit for superresolution via sparsity. Preprint at, https://arxiv.org/abs/1502.01385 (2015).
- Fannjiang, A.C. Compressive imaging of subwavelength structures. SIAM Journal on Imaging Sciences 2, 1277-1291 (2009).
- Aaron, J. & Chew, T.-L. A guide to accurate reporting in digital image processing–can anyone reproduce your quantitative analysis? Journal of Cell Science 134, jcs254151 (2021).




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in